SnapGene - Simply the best DNA manipulation software
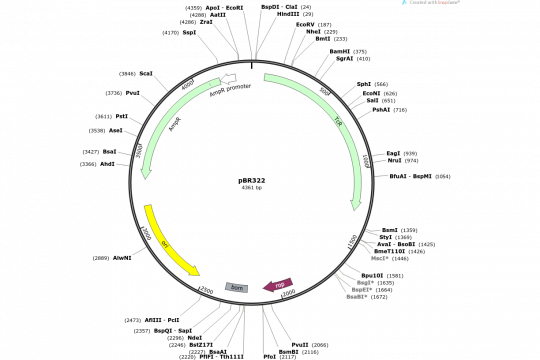
Our lab has been using different software packages to plan, document and visualize DNA constructs. Among those that we liked a lot for a long time were Textco's GeneConstructionKit (GCK) and Clone Manager (Professional). The latter runs unfortunately only under Windows. However, since several of our computers run Ubuntu Linux, we did run GCK versions 2.5 and 3 using WINE (a compatibility layer that allows us to run native Windows programs under Linux). However, with the upgrade to version 4, GCK became unusably slow under WINE and we were looking for a replacement. We contacted the developers of GCK, but they apparently were either not willing or able to help us. I suppose that the codebase of GCK is probably more than 20 years old and for that reason nobody dares to touch it. Just around that time, SnapGene was released and it fulfilled almost all of our requirements:
- True cross-platform: it runs on both Windows and Mac, and now even natively on Linux!
- It is able to open almost all other sequence formats. This was important for us as we had hundreds of our plasmid maps made with GeneConstructionKit. SnapGene reads among others: ApE, Clone Manager, DNASTAR Lasergene®, DNA Strider, EMBL (ENA), EnzymeX, GenBank/DDBJ, Gene Construction Kit®, MacVector, pDRAW32, Serial Cloner and Vector NTI®.
- It is not as expensive as the GeneConstructionKit or Clone Manager.
- It has a free version, the SnapGene Viewer, that everybody can download and use to view and print plasmid maps
- It is under active development: new features are added constantly. We have also put in one feature request: Virtual agarose gel images for partial restriction enzyme digests.
When we bought our first licenses, it was not yet natively available for Linux and hence we did run it under WINE, where it performed very well. We had some issues with opening files on remote samba shares, but it appeared that this was a problem of the gnome virtual file system when we mounted the shares via the Nautilus GUI (the bug still exists in Ubuntu 15.10; when you mount a samba share via the GUI, its mountpoint doesn't show up under /run/user/XXXX/gvfs and you need to force-unmount and manually start the gnome virtual file system again, after which the mountpoints magically appear: First, mount the samba share via GUI, then (via the command line): sudo umount -f /run/user/1000/gvfsz, then (as regular user) /usr/lib/gvfs/gvfsd-fuse /run/user/1000/gvfs, then mount the samba share again via GUI.
Even though they did not officially support SnapGene under WINE, the developers have helped us a lot with our WINE-related problems. When we organized a cloning course for the graduate school last year, the SnapGene guys arranged free licenses for all of the students for the duration of the course.
To top it all, with Version 3.0, SnapGene has recently released a native Linux port of SnapGene. We immediately bought additional licenses and have been installing the software on our Ubuntu machines (one 14.04 and three 15.10 machines). We also know that many other labs use and like this software, including Addgene (if anyone is in need of a good plasmid mapping software, it would be them).
For me, SnapGene is the way to go if you need a cloning software. What would be the next best thing if you can't pay for it? Probably SerialCloner. If you are an open source purist, you are out of luck: there are several free solutions, but none of them is open source...